Locality Sensitive Hashing
Locality Sensitive Hashing
Overview
Applications of set-similarity
- Many data mining problems can be expressed as finding
similarsets:- Pages with similar words, e.g., for classification by topic.
- Netflix users with similar tastes in movies, for recommendation systems.
- Dual: movies with similar sets of fans.
- Entity resolution.
- Example application: Given a body of documents, e.g., the Web, find pairs of documents with a lot of text in common, such as:
- Mirror sites, or approximate mirrors.
- Application: Don’t want to show both in a search.
- Plagiarism, including large quotations.
- Similar news articles at many news sites.
- Application: Cluster articles by
same story: topic modeling, trend identification.
- Application: Cluster articles by
- Mirror sites, or approximate mirrors.
Problem statement
- Given: High dimensional data points $x_1, x_2, …$
- For example: Image is a long vector of pixel colors
- And some distance function $d(x_1,x_2)$
- Which quantifies the
distancebetween $x_1$ and $x_2$
- Which quantifies the
- Goal: Find all pairs of data points $x_i, x_j$ that are within some distance threshold $d(x_i,x_j) \leq s$
- Naïve solution would take $O(N^2)$ where N is the number of data points
- This can be done in $O(N)$!!!
Motivation from frequent itemsets
- A-Priori:
- First pass: Find frequent singletons. For a pair to be a frequent pair candidate, its singletons have to be frequent!
- Second pass: Count only candidate pairs!
- PCY
- First pass: Count exact frequency of each item:
- Take pairs of items {i,j}, hash them into B buckets and count of the number of pairs that hashed to each bucket
- Second pass:
- For a pair {i,j} to be a candidate for a frequent pair, its singletons {i}, {j} have to be frequent and the pair has to hash to a frequent bucket!
- First pass: Count exact frequency of each item:
- What can we learn from PCY:
- First pass: Take documents and hash them to buckets such that documents that are similar hash to the same bucket
- Second pass: Only compare documents that are candidates (i.e., they hashed to a same bucket)
- Benefits: Instead of $O(N^2)$ comparisons, we need $O(N)$ comparisons to find similar documents
Finding Similar Items
Distance measures
- Goal: Find near-neighbors in high-dimensional space
- We formally define
near neighborsas points that are asmall distanceapart
- We formally define
- For each application, we first need to define what
distancemeans - Jaccard distance/similarity
- The
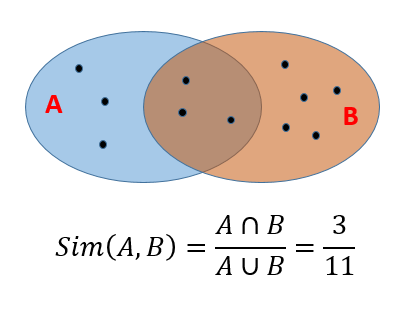
Jaccard similarityof twosetsis the size of their intersection divided by the size of their union:-
$sim(C_1, C_2) = C_1 \cap C_2 / C_1 \cup C_2 $
-
-
Jaccard distance:-
$d(C_1, C_2) = 1 - C_1 \cap C_2 / C_1 \cup C_2 $
-
- The
- The
Jaccard similiarityof two sets is the size of their intersection divided by the size of their union.
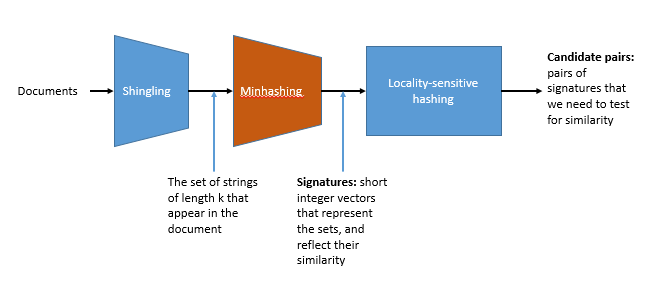
Three essential techniques for similar documents
- Shingling : convert documents, emails, etc., to sets.
- Minhashing : convert large sets to short signatures, while preserving similarity.
- Locality sensitive hashing : focus on pairs of signatures likely to be similar.
Shingling
Shingles
- Documents as high-dimensional data
- Simple approaches
- Document = set of words appearing in document
- Document = set of
importantwords - Don’t work well for this application: Need to account for ordering of words!
- Shingles
- A k-shingle (or k-gram) for a document is a sequence of k characters that appears in the document.
- Example:
- k = 2;
- Document:
abcab; - Set of 2-shingles: {ab, bc, ca}.
Compressing shingles
- To
compress long shingles, we can hash them to (say) 4 bytes - Represent a document by the set of hash values of its k-shingles
- Idea: Two documents could (rarely) appear to have shingles in common, when in fact only the hash-values were shared
- Example:
- k=2; document D1= abcab
- Set of 2-shingles: S(D1) = {ab, bc, ca}
- Select a random hash function $h$ such that
- Therefore, the hash of the shingles: h(D1) = {1, 5, 7}
Similarity metrics for shingles
- Document $D_i$ is a set of its k-shingles $C_i=S(D_i)$
- If we collect all unique k-shingles from all documents, each document can be represent as a 0/1 vector in the space of the k-shingles
- Each unique shingle is a dimension
- Vectors represent documents will be sparse, as no document should contain the majority of all k-shingles
Example
- Given the following documents:
- D1: practice makes perfect
- D2: perfect practice prevents poor performance
- D3: persistent practice produces progress
- The 1-shingle set for each documents are:
- D1: {practice,makes,perfect}
- D2: {perfect,practice,prevents,poor,performance}
- D3: {persistent,practice,produces,progress}
- The 1-shingle space:
- {makes, perfect, performance, persistent, poor, practice, prevents, produces, progress}
- The corresponding 0/1 vectors for each documents are:
- A natural similarity measure is the
Jaccard similarity
Motivation for Minhash/LSH
- Documents that are intuitively similar will have many shingles in common.
- k = 5 is ok for short documents
- k = 10 is better for long documents
- Problem: find near-duplicate documents among $N=10^6$ documents
- Naïvely, we would have to compute pairwise Jaccard similarities for every pair of docs
- $\frac{N(N-1)}{2} \approx 5 \times 10^{11}$ comparisons
- At $10^5\ secs/day$ and $10^6\ comparisons/sec$, it would take 5 days
- For $N=10^7$, it takes more than a year…
Minhashing: Jaccard Similarity
Encoding sets as bit vectors
- Many similarity problems can be formalized as finding subsets that have significant intersection
- Encode sets using 0/1 (bit, boolean) vectors
- One dimension per element in the universal set
- Interpret set intersection as bitwise AND, and set union as bitwise OR
- Example: C1 = 10111; C2 = 10011
- Size of intersection = 3; size of union = 4,
- Jaccard similarity (not distance) = 3/4
- Distance: $d(C_1,C_2)$ = 1 – (Jaccard similarity) = 1/4
Convert from sets to boolean matrices
- Rows: elements of the universal set. In other words, all elements in the union.
- Columns: individual sets.
- A cell value of
1in row e and column S if and only if e is a member of S. - A cell value of
0otherwise. - Column similarity is the Jaccard similarity of the sets of their rows that have the value of
1. - Typically sparse.
- This gives you another way to calculate similarity: column similarity = Jaccard similarity.
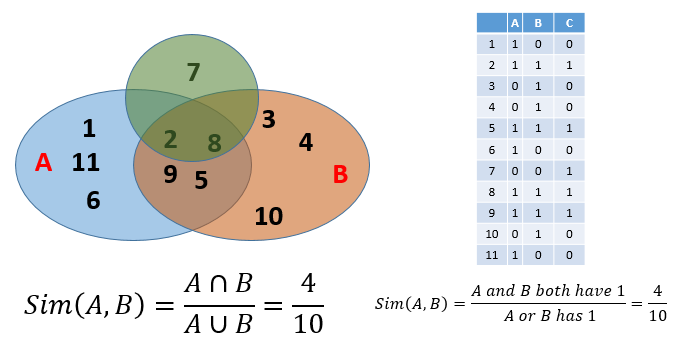
Finding similar columns
- Naïve approach:
- Signatures of columns: small summaries of columns
- Examine pairs of signatures to find similar columns
- Essential: Similarities of signatures and columns are related
- Optional: Check that columns with similar signatures are really similar
Hashing columns: Signatures
- Key idea: “hash” each column C to a small signature h(C), such that:
- (1) h(C) is small enough that the signature fits in RAM
- (2) $sim(C_1, C_2)$ is the same as the “similarity” of signatures $h(C_1)$ and $h(C_2)$
- Solution:
min-hashing
Minhashing
- Imagine the rows permuted randomly.
- Define
minhashfunction h(C) = the number of the first (in the permuted order) row in which column C has 1. - Use several (e.g., 100) independent hash functions to create a signature for each column.
- The signatures can be displayed in another matrix called the signature matrix
- The signature matrix has
- its columns represent the sets and
- the rows represent the
minhashvalues, in order for that column.
- Example: Assuming the following matrix for 4 documents, C1, C2, C3, and C4:
- 0/1 vector matrix:
- We permute the index of the row, then start going through the rows using the values in the
Permutationcolumn as indices.
Permutation 1
\[\left[ \begin{array}{ccccc} Permutation & C1 & C2 & C3 & C4 \\ 3 & 1 & 0 & 1 & 0 \\ 4 & 1 & 0 & 0 & 1 \\ 7 & 0 & 1 & 0 & 1 \\ 6 & 0 & 1 & 0 & 1 \\ 1 & 0 & 1 & 0 & 1 \\ 2 & 1 & 0 & 1 & 0 \\ 5 & 1 & 0 & 0 & 0 \\ \end{array} \right]\]
- For column C1:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 1 (first 1)
- Resulting signature is 2
- For column C2:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- For column C3:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 1 (first 1)
- Resulting signature is 2
- For column C4:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature set for this permutation is {2,1,2,1}
</details> <summary>Permutation 2</summary> [\left[ \begin{array}{ccccc} Permutation & C1 & C2 & C3 & C4
4 & 1 & 0 & 1 & 0
2 & 1 & 0 & 0 & 1
1 & 0 & 1 & 0 & 1
3 & 0 & 1 & 0 & 1
6 & 0 & 1 & 0 & 1
7 & 1 & 0 & 1 & 0
5 & 1 & 0 & 0 & 0
\end{array} \right]]
- For column C1:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 1 (first 1)
- Resulting signature is 2
- For column C2:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- For column C3:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 0
- row with value 3 in permutation column has a 0
- row with value 4 in permutation column has a 1 (first 1)
- Resulting signature is 4
- For column C4:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature set for this permutation is {2,1,4,1}
</details> <summary>Permutation 3</summary> [\left[ \begin{array}{ccccc} Permutation & C1 & C2 & C3 & C4
1 & 1 & 0 & 1 & 0
3 & 1 & 0 & 0 & 1
7 & 0 & 1 & 0 & 1
6 & 0 & 1 & 0 & 1
2 & 0 & 1 & 0 & 1
5 & 1 & 0 & 1 & 0
4 & 1 & 0 & 0 & 0
\end{array} \right]]
- For column C1:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- For column C2:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 1 (first 1)
- Resulting signature is 2
- For column C3:
- row with value 1 in permutation column has a 1 (first 1)
- Resulting signature is 1
- row with value 1 in permutation column has a 1 (first 1)
- For column C4:
- row with value 1 in permutation column has a 0
- row with value 2 in permutation column has a 1 (first 1)
- Resulting signature is 2
- Resulting signature set for this permutation is {1,2,1,2}
</details> - The resulting signature matrix is:
1
2
3
4
5
6
7
8
9
10
$$
\left[
\begin{array}{ccccc}
& C1 & C2 & C3 & C4 \\
1 & 2 & 1 & 2 & 1 \\
2 & 2 & 1 & 4 & 1 \\
3 & 1 & 2 & 1 & 2 \\
\end{array}
\right]
$$
Minhashing: surprising property
- The probability (over all permutations of the rows) that $h(C_1) = h(C_2)$ is the same as $Sim(C_1, C_2)$.
- The similarity of signatures is the fraction of the minhash functions in which they agree.
- The expected similarity of two signatures equals the Jaccard similarity of the columns.
- The longer the signatures, the smaller the expected error will be.
- Example:
- Given the following document matrix
- and its corresponding signature matrix
- The Jaccard similarity calculation for the two matrices are:
Minhashing: implementation
- Can’t realistically permute billion of rows:
- Too many permutation entries to store.
- Random access on big data (big no no).
- How to calculate hashes in sequential order?
- Pick approximately 100 hash functions (100 permutations).
- Go through the data set row by row.
- For each row r, for each hash function i,
- Maintain a variable M(i,c) which will maintain the smallest value value of $h_i(r)$ for which column c has 1 in row r.
Locality-sensitive-hashing (LSH)
Overview
- Generate from the collection of signatures a list of candidate pairs: pairs of elements where similarity must be evaluated.
- For signature matrices: hash columns to many buckets and make elements of the same bucket candidate pairs.
- Pick a similarity threshold
t, a fraction < 1.- We want a pair of columns
canddof the signature matrix M to be acandidate pairif and only if their signatures agree in at least fractiontof the rows.
- We want a pair of columns
LSH
- Big idea: hash columns of signature matrix M several times and arrange that only similar columns are likely to hash to the same bucket.
- Reality: we don’t need to study the entire column.
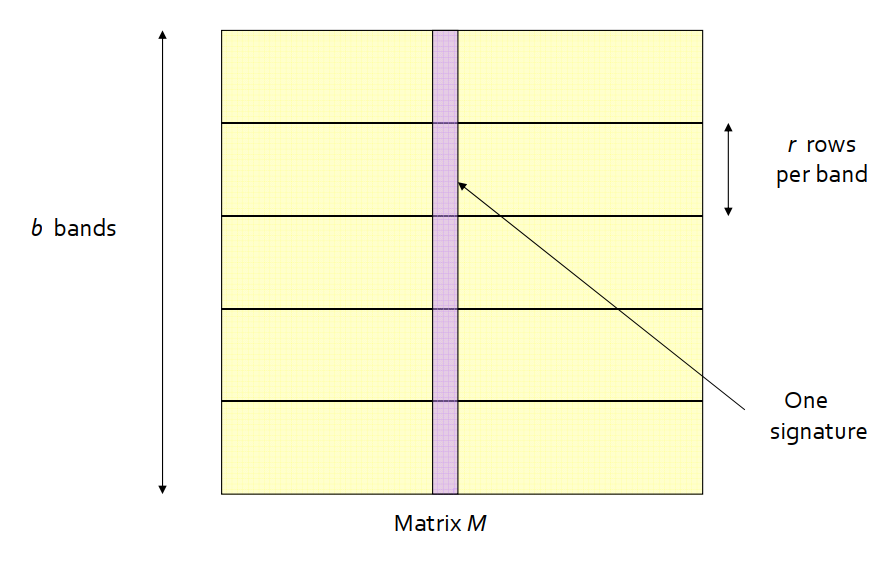
- Divide matrix M into
bbands ofrrows each. - For each band, hash its portion of each column to a hash table with
kbuckets, withkas large as possible. -
Candidatecolumn pairs are those that hash to the same bucket for at least one band. - Fine tune
bandr. - We will not go into the math here …
Hands on LSH
- Download the set of inaugural speeches from https://www.cs.wcupa.edu/lngo/data/inaugural_speeches.zip.